Chest X-ray Plus Artificial Intelligence Can Predict Worst Outcomes for COVID-19-Positive Patients
Computer program can accurately predict 80 percent of cases where COVID-19 patients will develop life-threatening conditions within four days.
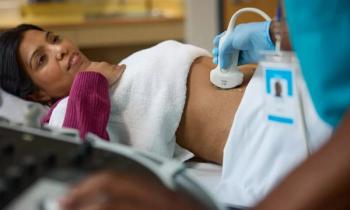
Chest X-rays aren’t just for diagnosing COVID-19. Newly published research now shows that these images can be part of a system that accurately predicts which patients will develop life-threatening conditions within four days.
By combining a deep neural network trained on chest X-rays with routine clinical variables, researchers from New York University Langone (NYU Langone) have created a system that performs comparably to interpreting radiologists and that can predict with 80-percent accuracy which COVID-19-positive patients will deteriorate, including mortality, intubation, and ICU admission.
“We develop[ed] an AI system that performs an automatic evaluation of deterioration risk, based on chest X-ray imaging, combined with other routinely collected non-imaging clinical variables,” said the team led by Krzysztof Geras, Ph.D., assistant professor of radiology at NYU Langone. “The goal is to provide support for critical clinical decision-making involving patients arriving at the [emergency department] in need of immediate care, based on the need for efficient patient triage.”
Related Content:
The team published details of their investigation on May 12 in
“Emergency room physicians and radiologists need effective tools like our program to quickly identify those COVID-19 patients whose condition is most likely to deteriorate quickly so that healthcare providers can monitor them more closely and intervene earlier,” said co-lead investigators Farah Shamout, Ph.D., assistant professor of computer engineering at New York University’s Abu Dhabi campus.
To create the neural network, the team used a training set of 5,224 chest X-rays from 2,943 patients who had severe COVID-19. With those images, they trained two models – one based on the Globally-Aware Multiple Instance Classifier (COVID-GMIC) and another based on a gradient boosting model (COVID-GBM).
Alongside details from the X-rays, the team incorporated patient age, race, and gender, as well as laboratory test results, such as weight, body temperature, and blood immune cell levels. They also factored in the need for a mechanical ventilator and whether the patient survived (2,405) or died (538).
Their test set consisted of 770 images from 718 patients who were admitted to NYU Langone between March 3, 2020, and June 28, 2020.
Related Content:
Based on their analysis, the team calculated the area under the receiver operating characteristic curve (AUC) and the area under the precision-recall curve (PR AUC) for deterioration prediction within 24, 48, 72, and 96 hours from the time of the chest X-ray. They, then, compared the system’s performance to outcomes produced by two radiologists.
Their main outcome, the team said, is that the deep neural network system outperforms the radiologists in terms of AUC and PR AUC at 48, 72, and 96 hours. Specifically, the COVID-GMIC achieved an AUC of 0.808 at 96 hours compared to 0.741 of both radiologists. This performance shows that the computer program accurately predicts four out of five cases where an infected patient will either require intensive care or mechanical ventilation or die within four days of admission.
“We hypothesize that COVID-GMIC outperforms radiologists on this task due to the currently limited clinical understanding of which pulmonary parenchymal patterns predict clinical deterioration, rather than the severity of lung involvement,” the team explained.
The team also modified the COVID-GMIC model to assess the probability that the first adverse event will happen within a certain time frame. At 96 hours, they said, the concordance index is 0.713, indicating that the modified model can effectively discriminate between patients.
“The system aims to provide clinicians with a quantitative estimate of the risk of deterioration, and how it is expected to evolve over time, in order to enable efficient triage and prioritization of patients at the high risk of deterioration,” the team explained. “The tool may be of particular interest for pandemic hotspots where triage at admission is critical to allocate limited resources, such as hospital beds.”
Geras’s team also sought to take their deep neural network one step further, pushing it to overcome the existing clinical implementation hurdle. On May 22, 2020, they silently deployed their system into the hospital where it was able to produce predictions in real-time in approximately two seconds.
Between the launch date and June 24, 2020, they collected 375 chest X-rays – 38 of which (10.1 percent) were associated with a positive 96-hour deterioration outcome. This outcome is lower than the 20.3 percent in the retrospective test set, but the differences are expected, the team said, due to potential patient differences and changing treatment guidelines as the pandemic progressed.
“The results suggest that the implementation of our AI system in the existing clinical workflows is feasible. Our model does not incur any overhead operational costs on data collection, since chest X-ray images are routinely collected from COVID-19 patients,” they said. “Additionally, the model can process the image efficiently in real-time, without requiring extensive computational resources, such as GPUs.”
But, there is still more work to be done, Geras said. Future work will examine whether employing a multi-modal strategy and including additional clinical data will improve performance. The team is also drafting clinical guidelines for the use of their classification test, and they hope to deploy it to emergency physicians and radiologists soon.